time_ordered_coords() is a helper function to create time-ordered DAGs.
Pass the results to the coords argument of dagify(). If .vars if not
specified, these coordinates will be determined automatically. If you want to
be specific, you can also use a list or data frame. The default is to assume
you want variables to go from left to right in order by time. Variables are
spread along the y-axis using a simple algorithm to stack them. You can also
work along the y-axis by setting direction = "y".
Arguments
- .vars
A list of character vectors, where each vector represents a single time period. Alternatively, a data frame where the first column is the variable name and the second column is the time period.
- time_points
A vector of time points. Default is
NULL, which creates a sequence from 1 to the number of variables.- direction
A character string indicating the axis along which the variables should be time-ordered. Either "x" or "y". Default is "x".
- auto_sort_direction
If
.varsisNULL: nodes will be placed as far"left"or"right"of in the graph as is reasonable. Default is right, meaning the nodes will be as close as possible in time to their descendants.- fixed_time
A named numeric vector pinning specific nodes to time points (e.g.,
c(x = 3, z = 1)). Only used in auto mode (.vars = NULL). Other nodes are placed automatically while respecting these constraints. Pinned times are 1-based and preserved in the output.- adjust_exposure_outcome
If
TRUE(default), automatically shift the outcome forward by one time point when it shares a layer with the exposure. All descendants of the outcome are also shifted. Only applies in auto mode and when the DAG has exposure and outcome set.- force_y
If
TRUE(default), run force-directed Y optimization to minimize node-edge overlaps. IfFALSE, nodes are evenly spaced within each layer using barycenter ordering only. Setting toFALSEis useful when edges will be curved or auto-routed, where tight Y positioning is less important. Only used in auto mode (.vars = NULL).
Examples
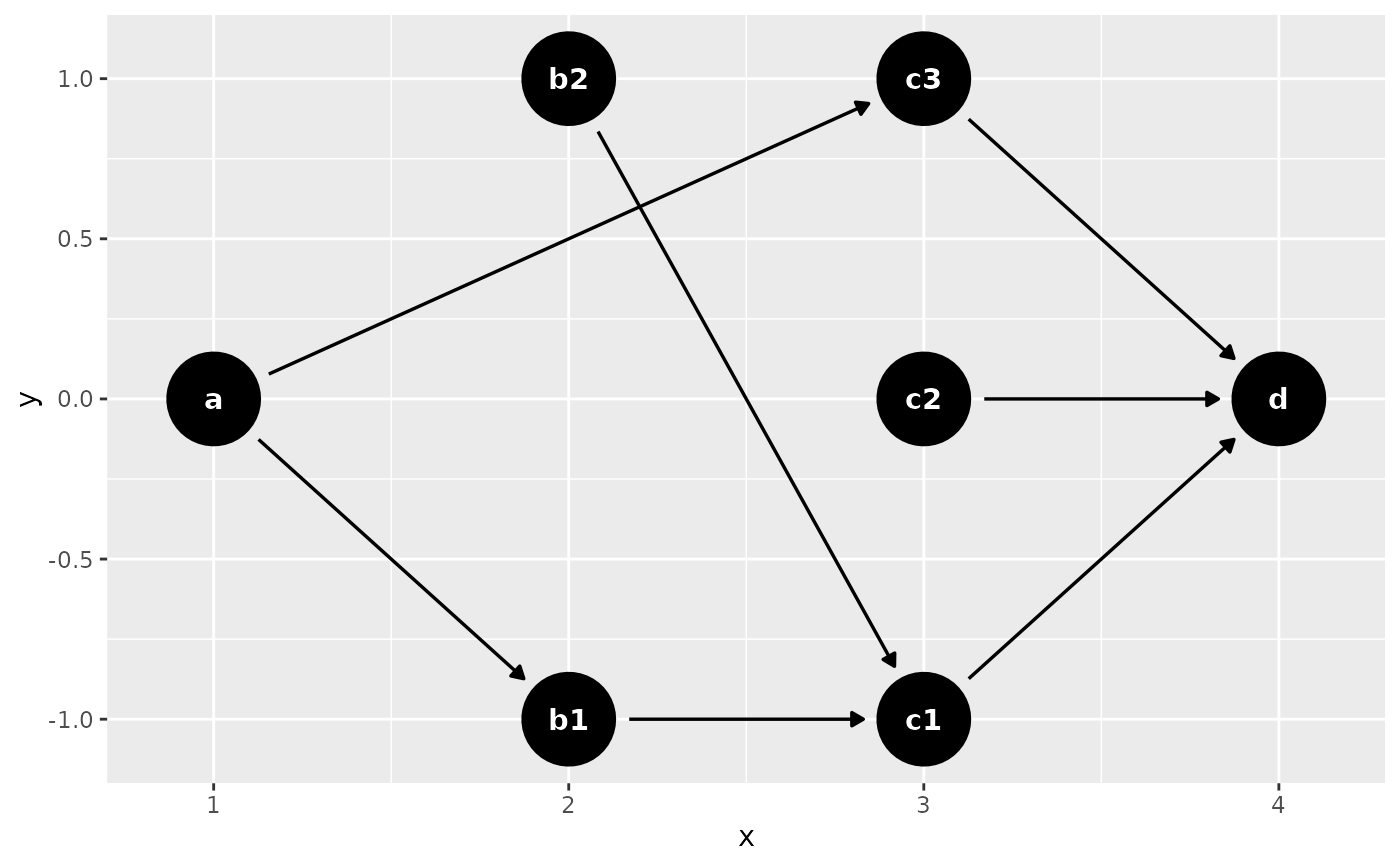
dagify(
d ~ c1 + c2 + c3,
c1 ~ b1 + b2,
c3 ~ a,
b1 ~ a,
coords = time_ordered_coords()
) |> ggdag()
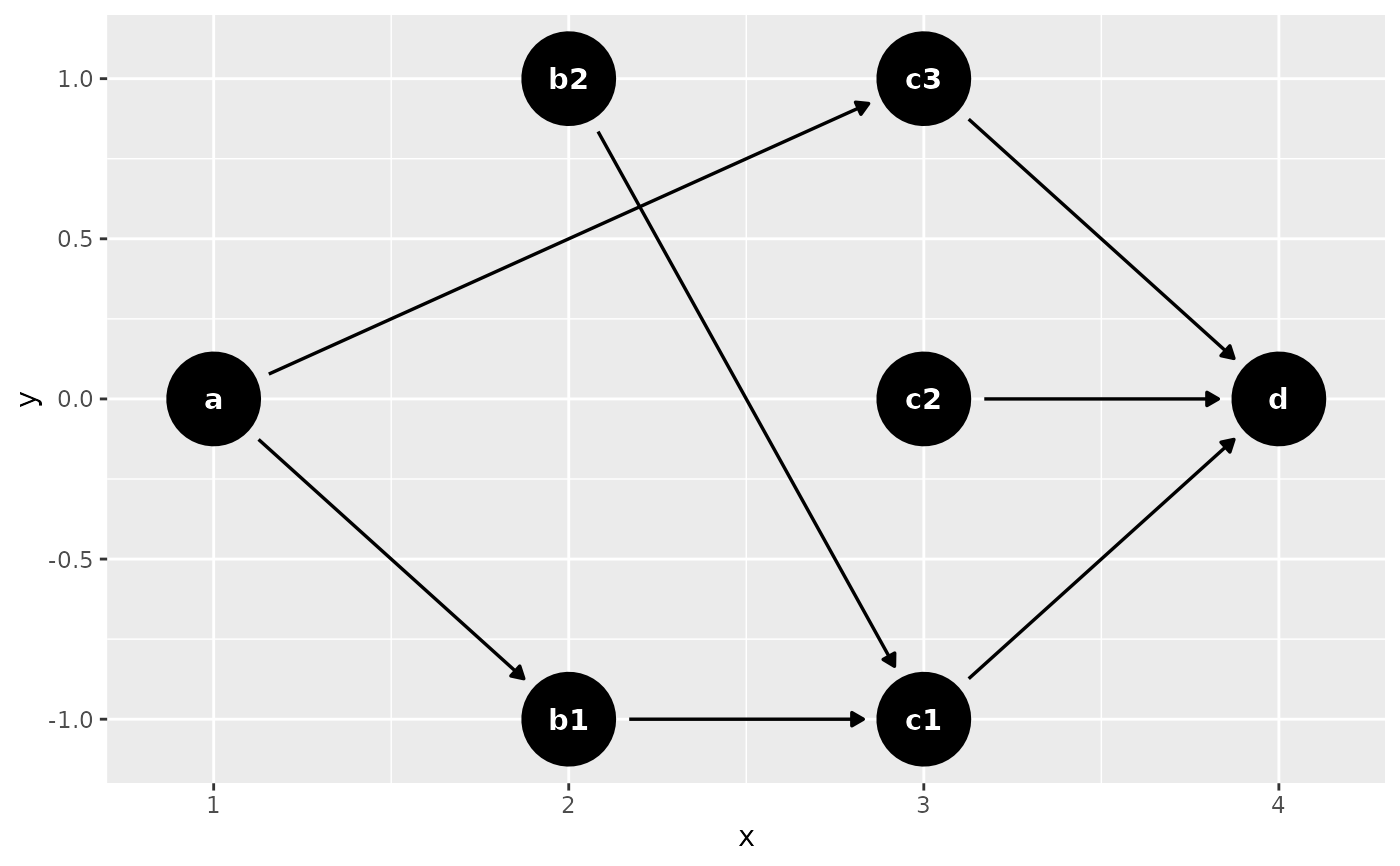
 coords <- time_ordered_coords(list(
# time point 1
"a",
# time point 2
c("b1", "b2"),
# time point 3
c("c1", "c2", "c3"),
# time point 4
"d"
))
dagify(
d ~ c1 + c2 + c3,
c1 ~ b1 + b2,
c3 ~ a,
b1 ~ a,
coords = coords
) |> ggdag()
coords <- time_ordered_coords(list(
# time point 1
"a",
# time point 2
c("b1", "b2"),
# time point 3
c("c1", "c2", "c3"),
# time point 4
"d"
))
dagify(
d ~ c1 + c2 + c3,
c1 ~ b1 + b2,
c3 ~ a,
b1 ~ a,
coords = coords
) |> ggdag()
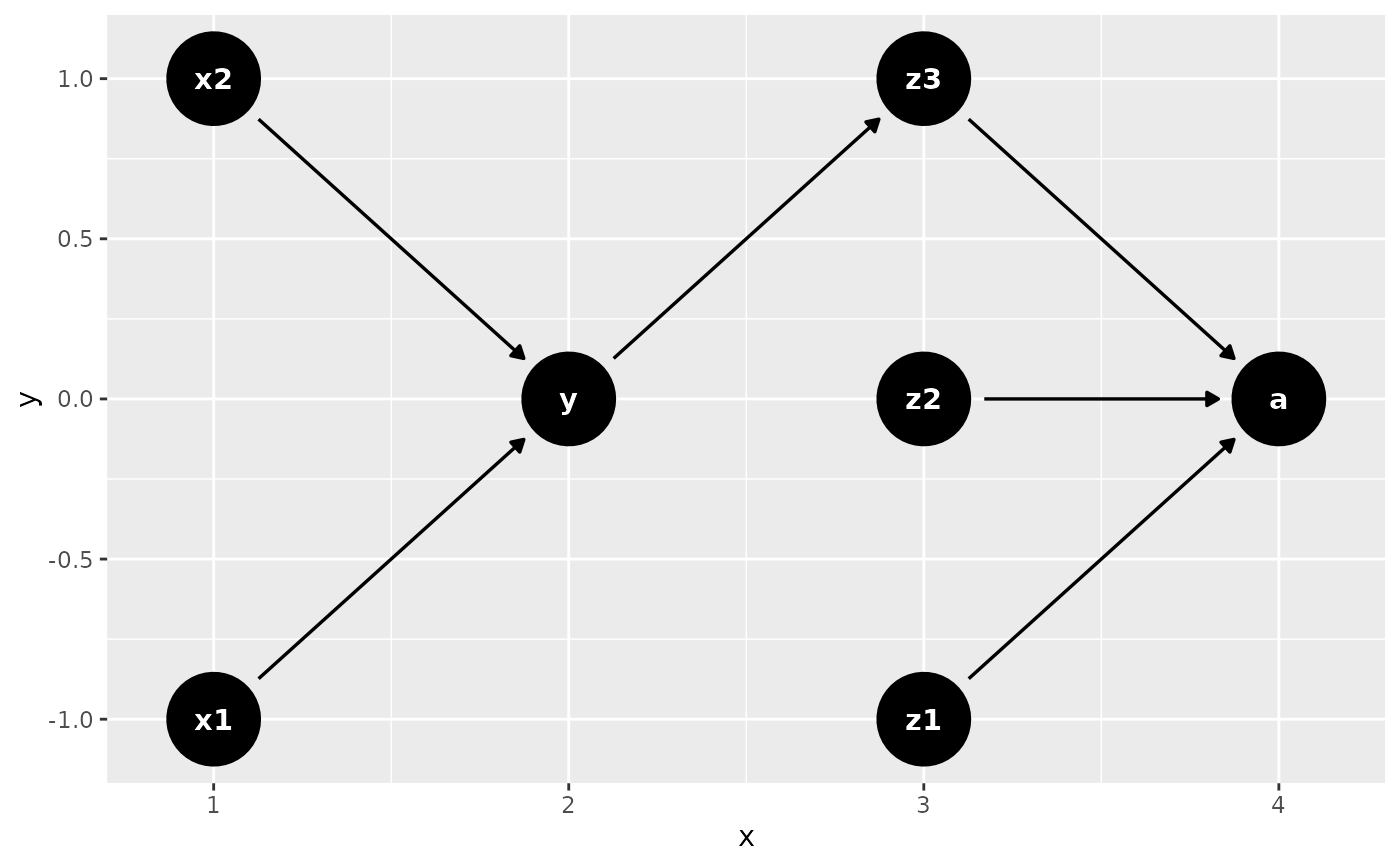
 # or use a data frame
x <- data.frame(
name = c("x1", "x2", "y", "z1", "z2", "z3", "a"),
time = c(1, 1, 2, 3, 3, 3, 4)
)
dagify(
z3 ~ y,
y ~ x1 + x2,
a ~ z1 + z2 + z3,
coords = time_ordered_coords(x)
) |>
ggdag()
# or use a data frame
x <- data.frame(
name = c("x1", "x2", "y", "z1", "z2", "z3", "a"),
time = c(1, 1, 2, 3, 3, 3, 4)
)
dagify(
z3 ~ y,
y ~ x1 + x2,
a ~ z1 + z2 + z3,
coords = time_ordered_coords(x)
) |>
ggdag()